Debbink Research Lab
Assistant Professor, Department of Natural Sciences
301-860-3341
kdebbink@bowiestate.edu
The Debbink Research Lab trains undergraduate researchers in microbiology and virology.
Projects
vein associated virus
Encapsidation of citrus yellow vein associated virus
The Debbink studies the functional relationships that dictate the ability of umbraviruses to be encapsidated by helper luteovirus capsids. Our specific work involves a novel independently mobile RNA related to umbraviruses that infects citrus plants (CYVaV). We are interested in the interaction between the CYVaV RNA and the coat protein of CVEV, its helper virus. Specifically, our goal is to map the encapsidation sequence that facilitates this interaction.
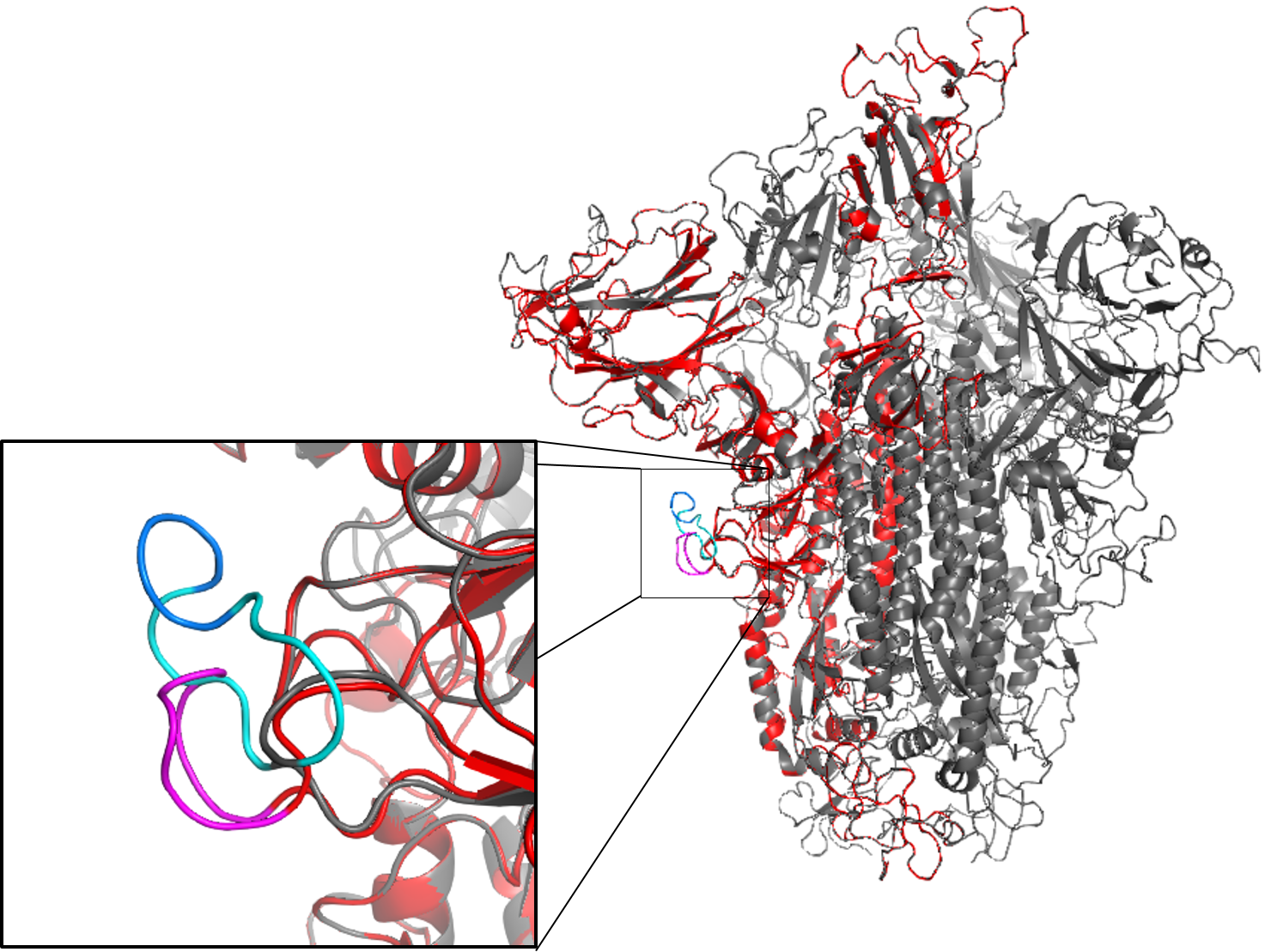
overlaid with a homology model of a deletion mutant
monomer (red) in the loop region.
Bioinformatics Analysis of Structure-Function Relationships for Viral Surface Proteins
The Debbink studies the interplay between structure and function of viral surface proteins for coronaviruses and noroviruses. We use computational programs to predict the impacts of different mutations on the function of the protein. Generally, we are interested in how changes in amino acid sequences coding for viral surface proteins impact the structural state of the virus, mapping antibody binding and neutralization sites on viral surface proteins, and detecting patterns in structure/function relationships among phylogenetically similar viruses. We collaborate with the Menachery Lab at UTMB on SARS-CoV2 studies.
Current Researchers
 Maame Ackon
Maame Ackon
 Steeven Tabuada
Steeven Tabuada
Alumni
- Bai Bangura
- Idris Gbadamosi
- Taylor Hailstock
- Jacquan Hilliard
- Collins Kibet
- Layelle Leach
- Daniel Ogunmiloro
- Whitney Okoe
- Steeven Tabuada
- Kammula Wesson
